Histology / Cell Biology Flashcards
(110 cards)
What is angiogenesis?
The growth of new blood vessels from preexsiting vessels
Tumor cells ___ ___invade normal tissue and capture existing blood vessels, rather tumor cells ________ normal endothelial cells to form new blood vessels.
Tumors DO NOT invade normal tissue and capture existing blood vessels, rather tumors recruit normal endothelial cells to form new blood vessels
Describe the general process of angiogenesis
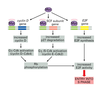
Cells typically monitor Oxygen (O2) tension. Under hypoxic conditions cells release angiogenic factors such as VEGF (vascular endothelial growth factors). VEFG binds to tyrosine kinase receptors on the surface of endothelial cells stimulating their proliferation. Endothelial cells migrate towards areas of higher VEGF concentration. In this way, VEGF can act as a chemoattract.
What is the angiogenic “switch”?
- Normal conditions prevent cells from triggering angiogensis
- As tumors progress they gain the capacity to promote angiogenesis
- Thus, overcoming the normal inhibition of angiogenesis is a step or switch, in tumor progression
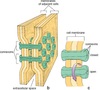
What are the differences between normal capillaries and tumor capillaries?
Tumor capillaries are:
- Larger - 3 times the diameter
- Disorganized layout
- Loosely associated pericytes
- Gaps of several microns between cells
- Permeable - Walls up to 10 times more permeable
- High Pressure - Higher interstitial fluid pressure
- Poor Drainage - Poor lymphatic drainage
Name some angiogenic growth factors
- VEGF (Vascular Endothelial Growth Factor)
- TGF-beta (Transforming Growth Factor Beta)
- bFGF (Basic Fibroblast Growth Factor)
- Interleukin-8
- Angiopoietin
- Angiogenin
- PDGF (Platelet Derived Growth Factor)
How do tumor cells induce angiogenesis
- Tumor cells can attract stromal cell types, which in turn can promote angiogenesis
- Tumor cells may release enzymes (like MMPs) which degrade ECM and the basement membrane
- Sometimes other angiogenic growth factors are present in ECM. When it is degraded those are released leading to additional angiogenesis.
- Tumor cells may attract local endothelial cells, but can also attract precursor cells from distant sites like the bone marrow.
Name some Anti-Angiogenic growth factors
Angiogenic Inhibitors
- Thrombospondin-1: ECM glycoprotein. Stops endothelial cells from proliferating
- Arrestin: fragment of collagen type IV
- Tumstatin: fragment of collagen type IV
- Angiostatin: fragment of plasminogen
- Endostatin: fragment of collagen XVIII
- Fibulin: fragment of basement membrane
- Endorepellin: fragment of perlecan
- TIMP-2: inhibitor of matrix metaloproteinase-2
What biological processes rely on angiogenesis?
- Embryonic development
- Implantation of the placenta
- Wound healing
- Disease processes
- Diabetic retinopathy, psoriasis, rheumatoid arthritis
- Tumorigenesis
What are the stages of the cell cycle? What are their relative lengths?
- S Phase (8-12 hours)
- M Phase (1 hour)
- G1+G2 (11-13 hours)
Interphase is G1+S+G2
Rapidly proliferating cells go through the cell cycle faster
Why are there checkpoint controls in the cell cycle?
Checkpoints provide an opportunity for a cell to determine if it is in a favorable condition to continue in the cell cycle. Is all the DNA replicated? Are all chromosomes attached to the spindle? Is the environment favorable?
Checkpoints allow for delays due to internal issues such as damaged DNA or poorly attached chromosomes
Checkpoints also allow for accumulation of microenvironmental signals such as growth factors indicating a general need to cell cycle progression in the tissue bed.
Where are checkpoints located in the cell cycle? What happens at each checkpoint?
(1) Before entering S phase from G1– Is the environment favorable? Checkpoint prevents S phase entry by stimulating p53 that activates transcription of the CKI called p21 that inhibits both G1/S-Cdk and S-Cdk
(2) Before entering M phase from G2– is all the DNA replicated? Checkpoint prevents M phase entry by stimulating kinases that inactivate the Cdc25 phosphatase needed for M-Cdk activation
(3) Before cytokinesis in M phase– are all the chromosomes attached to the spindle? Mad2 binds unattached kinetochores and blocks Cdc20-APC induced destruction of securin by sequestering Cdc20
How do checkpoints operate?
By NEGATIVE signals designed to stop the action
How were cell cycle control genes initially predicted and validated? (list experiments)
Genetic approach: Temperature sensitive yeat mutants– cell cycle control genes were predicted by G1/S blockade. Essential kinases are rate0limiting for cell cycle progression, and if these kinases are non functional, cell cycling stops at point where the kinases are needed.
Biochemical approach: Xenopus frog egg cytoplasm combined with the nuclei from a frog sperm and ATP– caused cell-free mitotic cycle, providing evidence for proteins in the cytoplasm driving mitosis and causing chromosomes to replicate and condense
Cell biology approach: Imaging replicated DNA in single cells or cell populations provides kinetic and spatial information about S phase activity– can see DNA replication occuring via an image, and can see relative amounts of DNA increasing during S phase
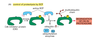
What are cyclins?
Cyclins are rate-limiting proteins that bind kinases (cyclin dependent kinases, or Cdk)
Cyclins undergo cycles of synthesis and degradation in each cycle
Cyclins and Cdk-activating kinase (CAK) are needed to activate Cdk
How is Cdk activity modulated?
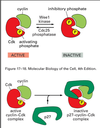
(1) Inhibitory phosphorylation, such as Wee1 kinase with M-Cdk. The Wee1 kinase adds an extra phosphate which causes the inactivation of M-Cdk. Cdc25 phosphatase removes the extra phosphate to activate M-Cdk
(2) Cyclin-dependent kinase inhibitor (CKI), an inhibitory protein which binds to the active Cdk and caused an inactive CKI-Cdk complex, such as with G1/S Cdk and p27 CKI
(3) Ubiquitin ligase, reserves of cyclin:Cdk complexes can also be deployed or destroyed through ubiquitinization
–Anaphase Promoting Complex (APC) ligates ubiquitin to M-cyclin to end M phase
–SCF adds ubiquitin to p27 CKI to activate G1/S Cdk
Can the cytosol from an S phase nucleus trigger DNA replication in the G1 or G2 nucleus?
Only in G1 nucleus
What are the 4 phases of the cell cycle and what happens during each one?
G1: gathering nutrients and synthesizing RNA and proteins necessary for DNA synthesis
S: DNA replication
G2: cell growth and reorganization of organelles
M: mitosis
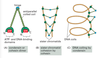
Describe the steps involved in replicating DNA before, during, and after S phase.
(1) Pre-replicative phase in G1: Cdc6 binds to the origin recognition complex (ORC) and MCM is recruited.
(2) S phase: S-Cdk phosphorylates Cdc6 causing its removal from the ORC and eventual degradation, MCM and phosphorylated ORC create replication fork.
(3) Upon the completion of DNA replication, M-Cdk phosphorylates MCM and Cdc6 creating a lack of the pre-replicative complex, so no more DNA can be replicated
Which cyclin-Cdk complex regulates G1 phase progression?
Cyclin D and Cdk4/6
What happens at the DNA replication checkpoint?
- Unfinished replication forks send a negative signal to M-Cdk
- Negative signal activates a kinase that inhibits the Cdc25 phosphatase and allows Wee1 to keep M-Cdk in inactive state
- M-Cdk is very important in driving mitosis, so inhibiting it will allow for replication forks to finish
What does M-Cdk cause?
M-Cdk induces assembly of mitotic spindle, chromosome condensation, nuclear envelope breakdown, actin-myosin cytokinesis, and distribution of membranous organelles to daughter cells
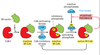
How are sister chromatids separated?
M-Cdk causes two actions:
(1) Promotes Cdc20 to bind to inactive APC (anaphase promoting complex) to make active APC–> active APC causes the degradation of securin and activation of separase
(2) Mitotic spindle formation and attachment to the sister chromatids, separase cleaves cohesins
Now chromosomes can move toward the poles
How does the spindle-attachment checkpoint work?
Mitosis
- Sensor detects whether the sister chromatids are connected to spindle aparatus and moving to opposite pole
- If there is a failure in this, it will send a negative signal to block anaphase
- Mad2 binds unattached kinetochores and blocks Cdc20-APC induced destruction of securin by sequestering Cdc20
- M-Cdk and Polo kinase both release Cdc20 when all kinetochores become attached to the spindle, allowing for anaphase to proceed