Unit 1 - Signal Transduction Flashcards
(117 cards)
Molecular
activation of enzyme, generation/synthesis of metabolite and its degradation
cellular
number of mitochondria regulated in a cell may divide, split, fuse
tissue/organ
no of cells or cell types are tightly regulated
4 levels of regulation in living organism
molecular
cellular
tissue/organ
organism (neuro-endocrine)
main mechanims of integration of the levels of regulation
HPA axis

what is the target of regulation

definition of signal transduction
process linking the signal-activated receptor and the biological response
there is a common logic in the structure of signal transducers and in the mode of their function
how is regulation initiated and and how does it flow
initiated by the signal (information) reaching the cell or formed within the cell

EC signal molecule → receptor protein → IC signalling proteins → target proteins

what can a signal be, based on their origin
from environment - pin, temp, light, smell
from organism - hormones, cytokines, metabolites
from within the same cell - DNA damage, ROS
what can a signal be if chemical
- proteins - GH
- peptides - insulin, neuropeptides
- AAs, derivatives - thyroxine, dopamine, epinephrine
- lipids - PGs, platelet activating factor
- ions - Ca2+, Cl-
- nucleotides - adenosines with specialised receptors on their surface
- gases - nitric oxide (produced by endothelial cells – trigger smooth muscle relaxation and vasodilation)
types of signal transmission

what is the significance of ligand binding
specificity
amplification
co-ordination of response - the same receptor may be present on a number of cells
cell specific response - by regulating the number and function of receptors
example of disease that involves a cell-specific response
type 2 diabetes
autoinhibitory receptors are present on cells become inactivated and unresponsive
types of receptors
cell surface/transmembrane receptors
nuclear receptors
cytosolic receptors

receptors for what hormone are present on a wide variety of cells
cortisol
amplification system
Uniform response of cell
e.g. If a cell triggers lipid degradation, whole lipid synthesis across the cell has to shut down and lipid degradation must occur
Net 0 effect - what 1 process achieves, the other will undo

allosteric regulation

interconversion cycle
NOTE: enzyme specific - some are active phosphorylated, some are active dephosphorylated
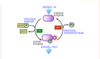
Akt/Protein Kinase P interconversion cycle - 1st mechanism
inactive Akt → Akt by 2 mechanisms:
FIRST
phosphorylated serine 473 interacts with the linker between the kinase and pH domains

2nd mechanism in Akt/protein kinase P interconversion cycle
phosphorylated serine 477 and threonine 479 result in the displacement of the pH domains

enzyme cascade

acceptors
all signal transduction pathways culminate to control of function of proteins (mostly enzymes, ion channels, transporters), which are involved directly in formation of biological response
2 categories of acceptors
control of protein amount
gene expression
protein degradation
protein stabilisation (tumour suppression gene P53)
control of existing proteins
without covalent modification of the protein (P) - Allosteric activator binds to an allosteric enzyme complex
with covalent modification of the protein - reversible and irreversible (cleavage of 3 forms of enzymes e.g. digestive enzymes involved in cleavage of trypsinogen to inter trypsin)




































