Will a 4 or 6 B.P sequence occur more frequently?
Obviously, by simple mathematical calculations, a 4 B.p sequence is more probable to occur than a 6 B.p sequence.
4 B.p - 44 –> 256 B.p
6 B.p - 46 –> 4096 B.p
Hence, on avaerge, an endonuclease that recognizes 4 B.p will produce fragments with an average of 256 B.p whereas, an endonuclease that recognizes 6 B.p will produce fragments with an average length of 4000 B.p.
What are heteroschizomers?
Restriction enzymes that recognize different sites in an identical sequence.
What are isoschizomers?
Restriction enzymes that cut at the same site within the same sequence.
Do many enzymes make staggered cuts?
YIEShhhhh
Many enzymes make staggered cuts in DNA –> leaves single-stranded overhangs (known as sticky ends) –> this makes it easier to stitch together DNA.
The example below – EcoRI –> creates a 4 B.p overhang with a free 5’ phosphate end.

Why do enzymes create staggered cuts?
Restriction enzymes make staggered cuts because they recognise sequences that display twofold symmetry. Meaning that they are identical after rotating them 180 degrees.
Sequences with two-fold symmetry are known as palindromes.
Hence, in a way, the sequences read the same backwards and forwards –> remember to read the top strand left to right and the bottom strand right to left.
In this case –> EcoRI cuts between G and A on the top strand and between G and A on the bottom.

Why do restriction endonucleases not cut up their own host’s DNA?
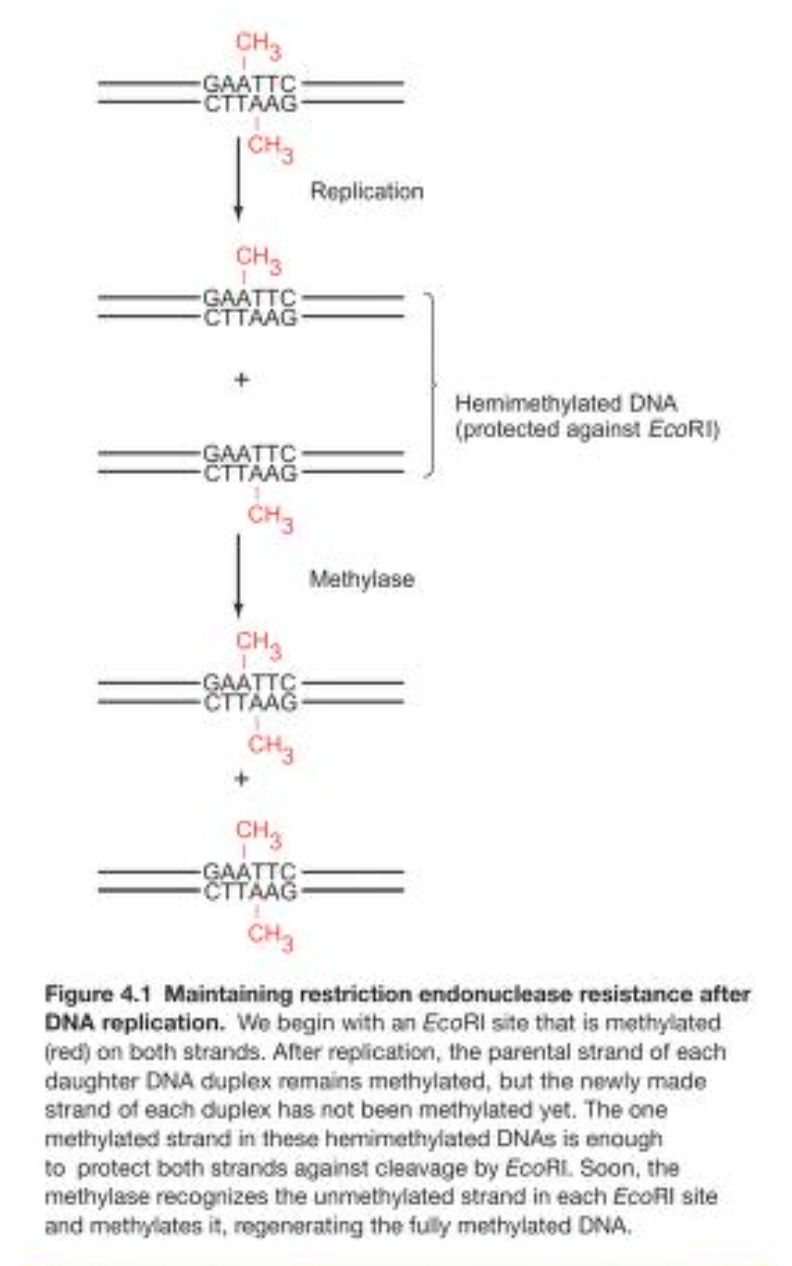
Almost all restriction endonucleases are paired with methylases that recognise and methylate the same DNA site as the restriction endonuclease.
Collective term for both is a restriction-modification system (R-M system).
After methylation the DNA sequence is protected –> will not be digested.
What happens during DNA replication, will the newly created unmethylated DNA not be cleaved?
Every time DNA replication takes place –> the daughter strands will have one new strand that is NOT methylated.
However, the other parental strand will be methylated.
This half-methylation is enough to protect the DNA against cleavage by most restriction endonucleases –> so this gives time for the methylases to methylate the new daughter strand –> yielding a fully methylated strand.
Things to consider when performing restriction mapping?
- Adding up all the fragments created by a single digest = total fragment size –> don’t add up? –> likely that fragments of the same size (i..e 2x 50 Kbp fragments) are superimposed on the gel.
- Examining the double digest —> multiple restriction enzymes digest DNA –> identify the common fragments and the unique fragments –> unique fragments are created by two restriction enzymes cutting a fragment.
Rough guide:
- Look at the first enzyme –> place a first restriction site at 12 O’clock –> place any other restriction site of this enzyme to match fragment sizes.
Check –> Are there any fragments retained in the double digest? If yes, then this fragment remains undigested by the other enzyme –> other regions will be digested instead.
- Then by trial and error try to place the restriction sites for the other enzyme ensuring that it matches the fragments produced by the single and double diegest.
What is southern and northern blotting used for?
Southern and northern blots are used to separate and identify DNA and RNA sequences, respectively.
Definition of restriction enzymes?
Restriction enzymes, also called restriction endonucleases –> recognize specific base sequences in double-helical DNA and cleave both strands of that duplex at the places
Where are restriction enzymes normally found?
Restriction enzymes are found in a wide variety of prokaryotes.
Their biological role is to cleave foreign DNA molecules (bacteriophages), providing the host organism with a primitive immune system.
Normally speaking, how long is the sequence that a restriction enzyme recognizes?
Many restriction enzymes recognize specific sequences of four to eight base pairs and hydrolyze a phosphodiester bond in each strand in this region.
How are restriction enzymes named?
Their names consist of a three-letter abbreviation for the host organ- ism (e.g., Eco for Escherichia coli, Hin for Haemophilus influenzae, Hae for Haemophilus aegyptius)
Followed by a strain designation (if needed)
Lastly, a roman numeral (to distinguish multiple enzymes from the same strain)
EcoRI
E. Coli strain R –> enzyme 1.
How can restriction fragments be separated and visualized?
Separation?
Using electrophoresis –> possible as phosphodiester backbone is negatively charged –> hence, DNA molecules will move when an electric field is applied.
Shorter –> further migration (visa-versa)
Visualization?
- Bands/spots of radioactive DNA in gels can be visualized by autoradiography.
- The gel can be stained with ethidium bromide –> fluoresces an intense orange under UV irradiation when double-helical DNA molecule is bound.
What are agarose gel used for? What are polyacrylamide gels used for?
Polyacrylamide gels –> Fragments with as many as 1000 bases
Agarose (more porous) –> larger fragments (as large as 20Kb)
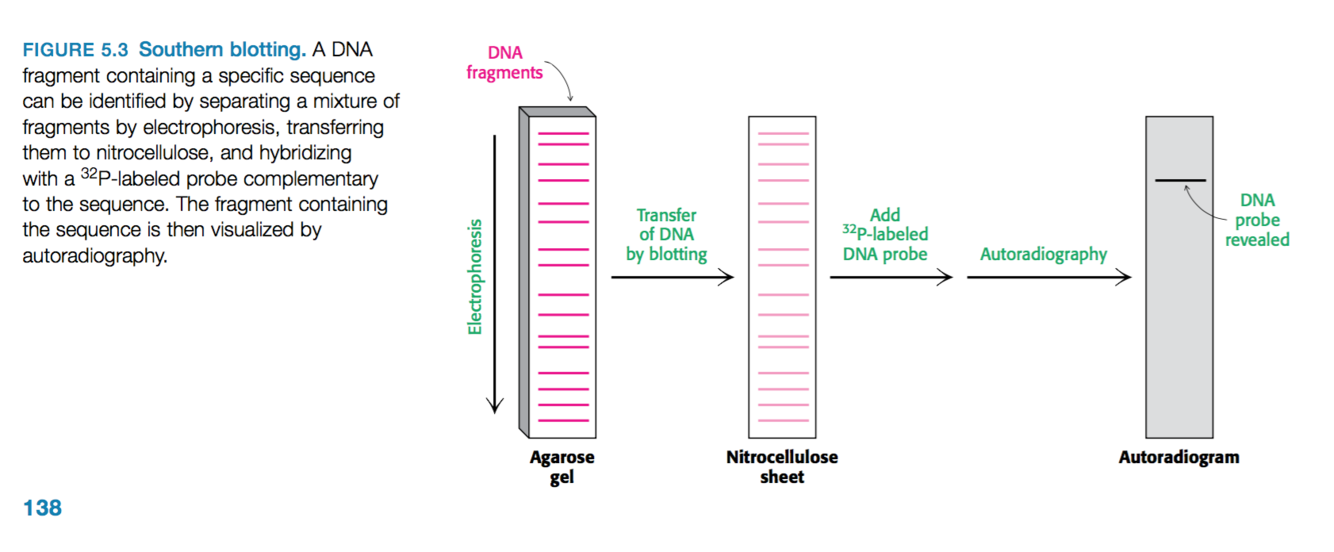
Outline the process of southern blotting.
Used to identify the presence of a particular base sequence in the sample of DNA.
- DNA is digested into fragments
- Restriction fragments undergo electrophoresis on an agarose gel.
- Denatured to form single-stranded DNA
- Transferred to a nitrocellulose sheet (polymer sheet) –> note the position of the DNA is preserved during this transfer event.
- Nitrocellulose sheet is then exposed to 32P-labeled or fluorescently tagged DNA probe –> short stretch of single-stranded DNA with a known base sequence.
- This probe will hybridize with a restriction fragment that has a complementary sequence.
- Then using autoradiography or fluorescence imaging we can find the position of the restriction-fragment–probe duplex
How does northern blotting differ from southern blotting?
Northern blotting is performed using a similar method as southern blotting but instead RNA molecules of a specific sequence can also be readily identified.
Sample procedure using electrophoresis, transfer to nitrocellulose and a DNA probe.
Outline the process of Sanger sequencing.
In a reaction vessel you have…
- Single-stranded DNA which you want the sequence for
- Primers
- DNA polymerase
- Deoxynucleotides
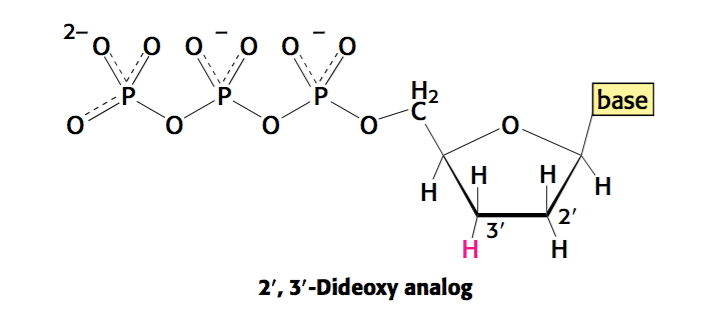
- A small quantity of di-deoxynucleotides of each nucleotide (each base contains a particular colour tag –> A is green, T is red, etc.)
Hence….
During the extension of the single strand –> if the analogue is incorporated (di-deoxy) –> replication will be terminated as the nucleotide lacks a 3 ‘ -OH to perform a nucleophilic attack.
Note –> di-deoxy is low enough that chain termination will take place occasionally.
Thus, in different reaction vessels, we have dNTPs and one particular di-deoxynucleotides –> i.e. analogue of dATP will terminate extension at every single T location –> thus, all the different chain lengths produced will correspond to the T position.
This process can be repeated for each type of nucleotide in order to deduce the sequence.
In the modern form of Sanger sequencing, how is the sequence readout?
Fragments are separated by a technique known as capillary electrophoresis.
This involves passing the mixture of DNA through a very narrow tube a gel matrix at high voltage (achieve separation quickly).
As they emerge from the tube –> we have a detector that detects the fluorescence given off by each base in the sequence –> gives us the base sequence very quickly.
What is needed for PCR?
DNA contains flanking regions that surround the target that we want to amplify.
- Primers that hybridize with the flanking sequences of the target.
- All four deoxyribonucleoside triphosphates (dNTPs)
- Heat-stable DNA polymerase –> Taq polymerase
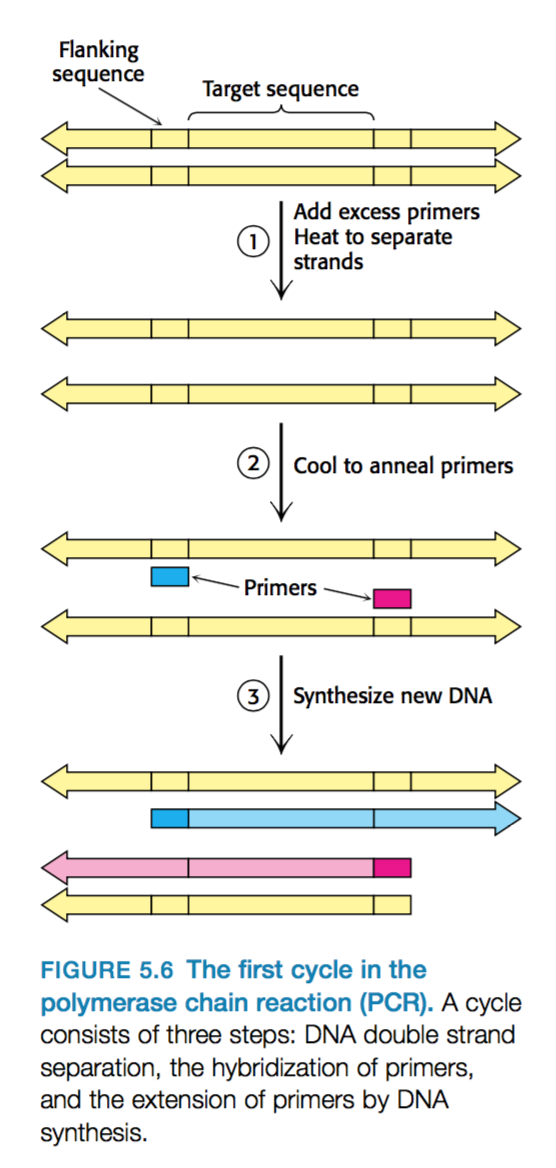
What are the three stages during PCR?
- Strand separation –> The two strands of the parent DNA molecule are separated by heating the solution to 95oC for 15s.
- Hybridization of primers (20-30 bases long) –> Solution cooled 54oC to allow the primer to hybridize to the DNA –> Primer anneals to 3’ region of one parental strand and the other primer also anneals to the 3’ region of the other parental strand.
- Parental strands do NOT rejoin because there is excess primers. - DNA synthesis –> Solution is heated to 72oC –> optimal temperature for taq which can then extend both strands in a 3’ to 5’ direction.
- Note –> polymerase extends further than target region.
This is one cycle which can be repeated over and over again to amplify DNA.
How to calculate the number of strands in PCR after ‘n’ cycles?
After ‘n’ cycles, the desired sequence is amplified 2n time.
Four advantages of running PCR?
One can amplify large amounts of DNA…
- Without knowing the target sequence
- The target can be much larger than the primers –> 10kb fragments have been amplified before.
- Primers don’t have to match flanking regions completely.
- It is highly specific due to the stringency of hybridization at a relatively high temperature (only the primer will anneal –> DNA with hybridized primers will only be extended)
Note - Stringency is the required closeness of the match between primer and target, which can be controlled by temperature and salt
High stringency (High temp/low salt) –> close match will anneal.
- PCR is VERY sensitive
Application of PCR?
Used as a diagnostics tool
- We can detect the presence of bacteria/viruses in a sample using a specific primer –> i.e. HIV.
Alternatively…
Can be used to detect the presence of cancer –> identify mutations of certain growth-control genes, such as the ras genes.
Furthermore…
In forensics and legal medicine –> amplify DNA samples from crime scenes.



